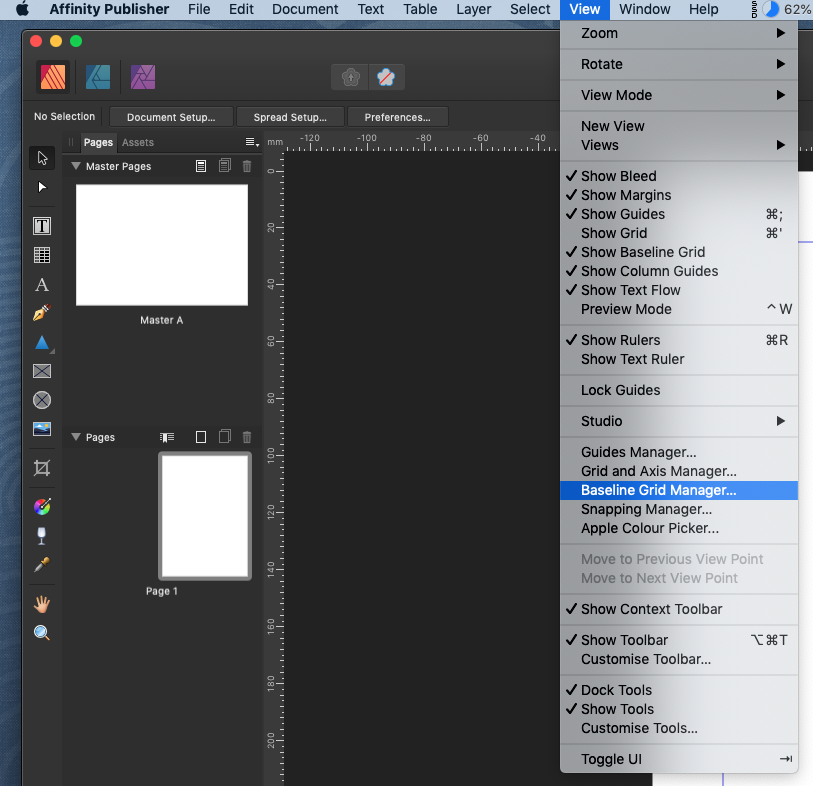
Mutation of these residues to alanine (A) disrupted ORF2p/PCNA copurification, whereas mutation of Q408, less conserved in ORF2p, does not affect it ( Figure 6B). Within the PIP sequence, four amino acid residues (I407, I411, Y414, and Y415) are highly conserved across L1 ORF2s from diverse species, suggesting a crucial role for a PIP box in ORF2 binding to PCNA. We identified a canonical PCNA-interacting protein (PIP) box (Qxxxx) in residues 407–415 of ORF2p, located between the EN and RT domains ( Figure 6A). 
PCNA was found as a low-abundance but high-specificity interactor of ORF2p, but not ORF1p. The second experiment was further optimized with respect to binding time and lysate to Dynabead ratios. 
LD288.1, LD288.2, LD401.1, and LD401.2 notation represent our first and second I-DIRT experiments, respectively, with tagged ORF1p and ORF2p, respectively. An arbitrary cutoff of > 71% heavy was assigned. ‡ Because the distribution is non-normal for the anti-ORF1p pullout for L1RP ORF1p-Flag, no p-Values can be generated and the %heavy is not normalized. Note that due to binning, the leftmost blue bar may contain some non-significant proteins only significant proteins are listed. P-values were determined by (see Experimental Procedures). Yellow bars represent non-significant groups, blue bars represent statistically significant specificity (i.e., percentage of heavy isotope content) after Benjamini-Hochberg correction for multiple hypothesis testing. Histograms plot the number of recovered proteins as compared to their heavy isotope content for six individual I-DIRT experiments. Images all taken using an epifluorescent microscope. In the right panels, within each cell type, the same exposure and normalization was used for each particular antibody. No cells were found to express ORF2p alone, except for pLD561, in which no cells were found to express ORF1p. 
In the left panels, images were captured using a 40x objective and normalized identically for each construct cells expressing ORF1p alone or both ORF1p and ORF2p at levels over background were counted. For Tet-On HEK293T LD cells, fibronectin-coated coverslips were used. Then puro resistant cells were plated on coverslips and induced by adding Dox 8-16 hr after plating. For remaining constructs with a pTRE promoter, HeLa or Tet-On HEK293T LD cells were transfected and puro selected for three days. 24 hr later, cells were fixed, permeabilized, and stained with anti-Flag and anti-ORF1 antibodies and then fluorescent-conjugated secondary antibodies. For constructs with a CMV promoter and pLD288, HeLa or Tet-On HEK293T LD cells were plated on coverslips and transfected 8-16 hr after plating. Graphical AbstractĪnti-ORF1 antibody used in all panels is rabbit monoclonal JH73. PCNA interacts with ORF2p via a PIP box motif mechanistic studies suggest that this occurs during or immediately after target-primed reverse transcription. Among the findings, UPF1, a key nonsense-mediated decay factor, and PCNA, the polymerase-delta-associated sliding DNA clamp, were identified and validated. These data sets include known interactors PABPC1 and MOV10 and, with in-cell imaging studies, suggest existence of at least three types of compositionally and functionally distinct L1 RNPs. Here, we describe a system to express and purify highly active L1 RNP complexes from human suspension cell culture and characterize the copurified proteome, identifying 37 high-confidence candidate interactors. As such, endogenous L1 expression levels are extremely low, creating a roadblock for detailed interactomic analyses. These streamlined elements require host factors to complete their life cycles, whereas hosts have developed mechanisms to combat retrotransposition’s mutagenic effects. LINE-1s are active human DNA parasites that are agents of genome dynamics in evolution and disease.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
